Towards genome editing cure of genetic heart diseases
10 years after CRISPR things are really heating up
This week 2 independent reports (here and here) demonstrated the potential for single-dose genome editing to approach hypertrophic cardiomyopathy (HCM), the most common form of heritable heart disease affecting 1 of 500 people. Prof Alanna Strong, the accompanying editorialist, while acknowledging many uncertainties, called for the beginning of Phase 1 clinical trials for genome editing of HCM.
Until recently, the idea that we could approach the heart with genome editing was not high on the list owing to formidable obstacles in delivery. Genome editing has achieved traction in clinical trials when the cells are directly accessible, such as editing T cells out of the body and administering them for cancer, or direct injection for treating blood or eye diseases, or targeting the liver because of its natural clearance role of intravenous infusions.
Then a series of 3 important papers from Eric Olson’s lab at UT Southwestern have changed the outlook. The first report last November targeted dilated cardiomyopathy in a mouse model that was created with homozygous (2 copies) of RNA binding protein motif 20 (RBM20) mutations, and led to rescue of heart function. Then in January 2023 a second paper targeted the calmodulin-dependent protein kinase (CAMKIIδ) gene, a key driver of heart disease, and demonstrated that editing could prevent heart damage in mice subjected to limiting and restoring heart blood supply. This week the same lab published the potential of curing hypertrophic cardiomyopathy in a humanized mouse model. Independently, this week the Seidman lab at Harvard also published the ability to prevent HCM in a mouse model. How has this remarkable and accelerated progress been made possible?
The Expanded Editor Menu
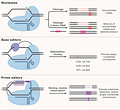
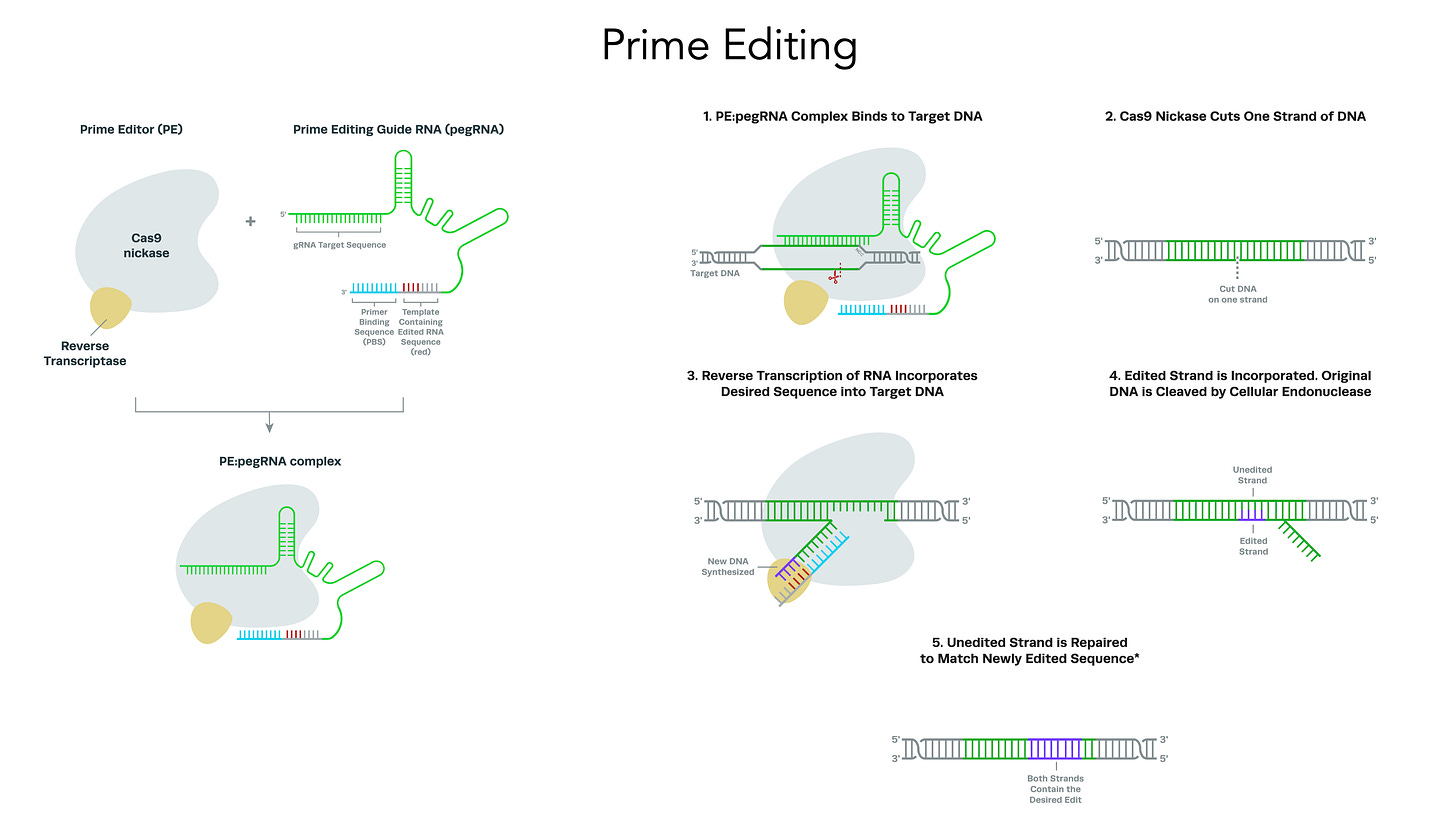
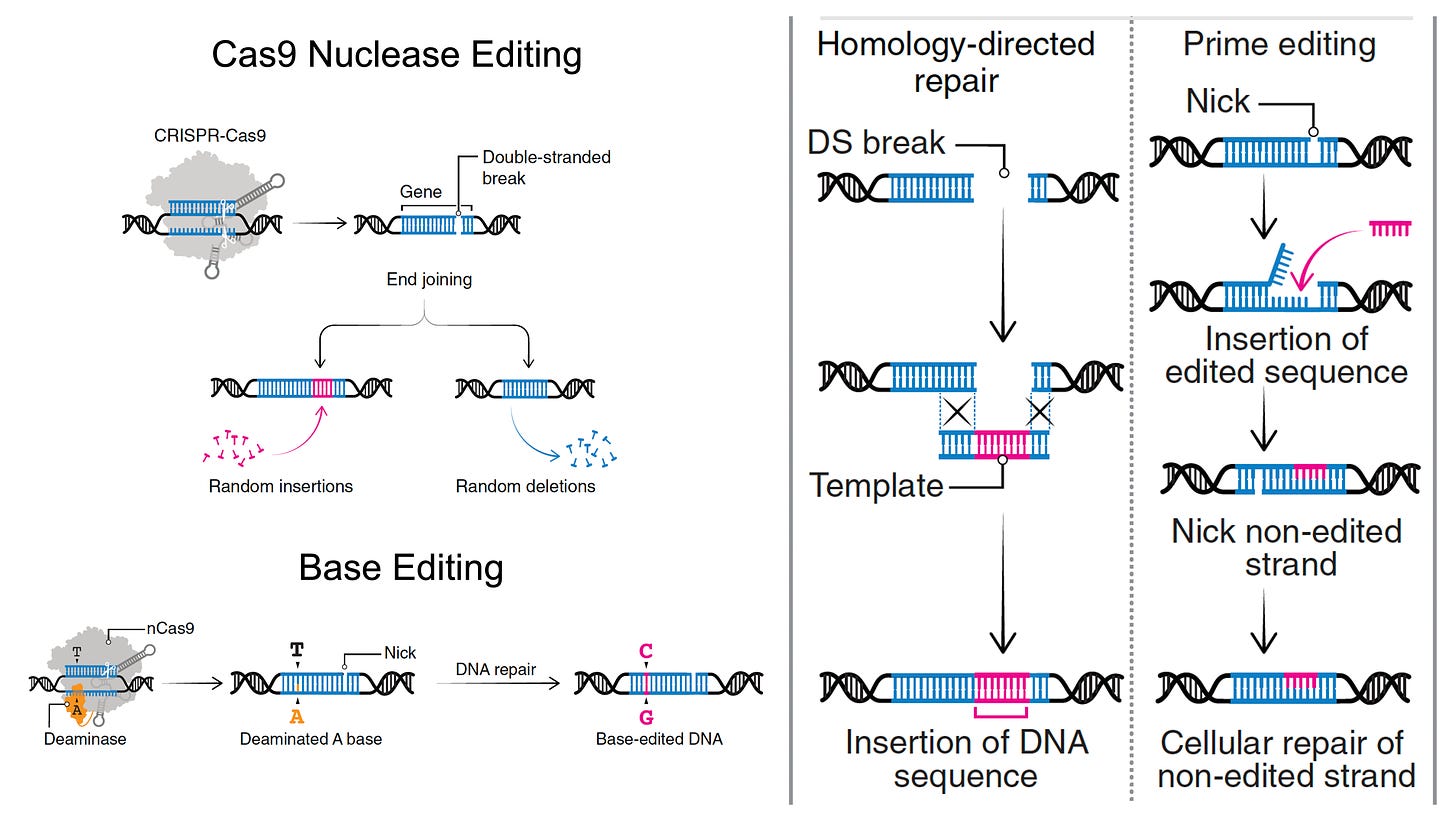
Most genetic diseases are due to a single nucleotide mutation. The original discovery of CRISPR for genome editing harnessed Cas9 nucleases that induce double-strand breaks in DNA, ideal for knocking out a disease-causing gene, but not suitable for affecting single nucleotide changes. That’s what base editors do, initially described by David Liu and colleagues in 2016, nicking the single-strand of DNA to affect the change (Liu described Adenine base editors (ABE) that change a A to a T, and there are Cytosine base editors (CBE) that change an C to a G. And subsequently came the invention of Prime editors (also from Liu and colleagues), a more complicated process involving reverse transcriptase, shown below, that sets up the potential for precise and more versatile genome manipulation, enabling the whole range of insertions, deletions and substitutions with single-strand DNA breaks and working in dividing and non-dividing cells. For those interested in more depth and future direction of Prime editing, there’s a recent and excellent review paper here.
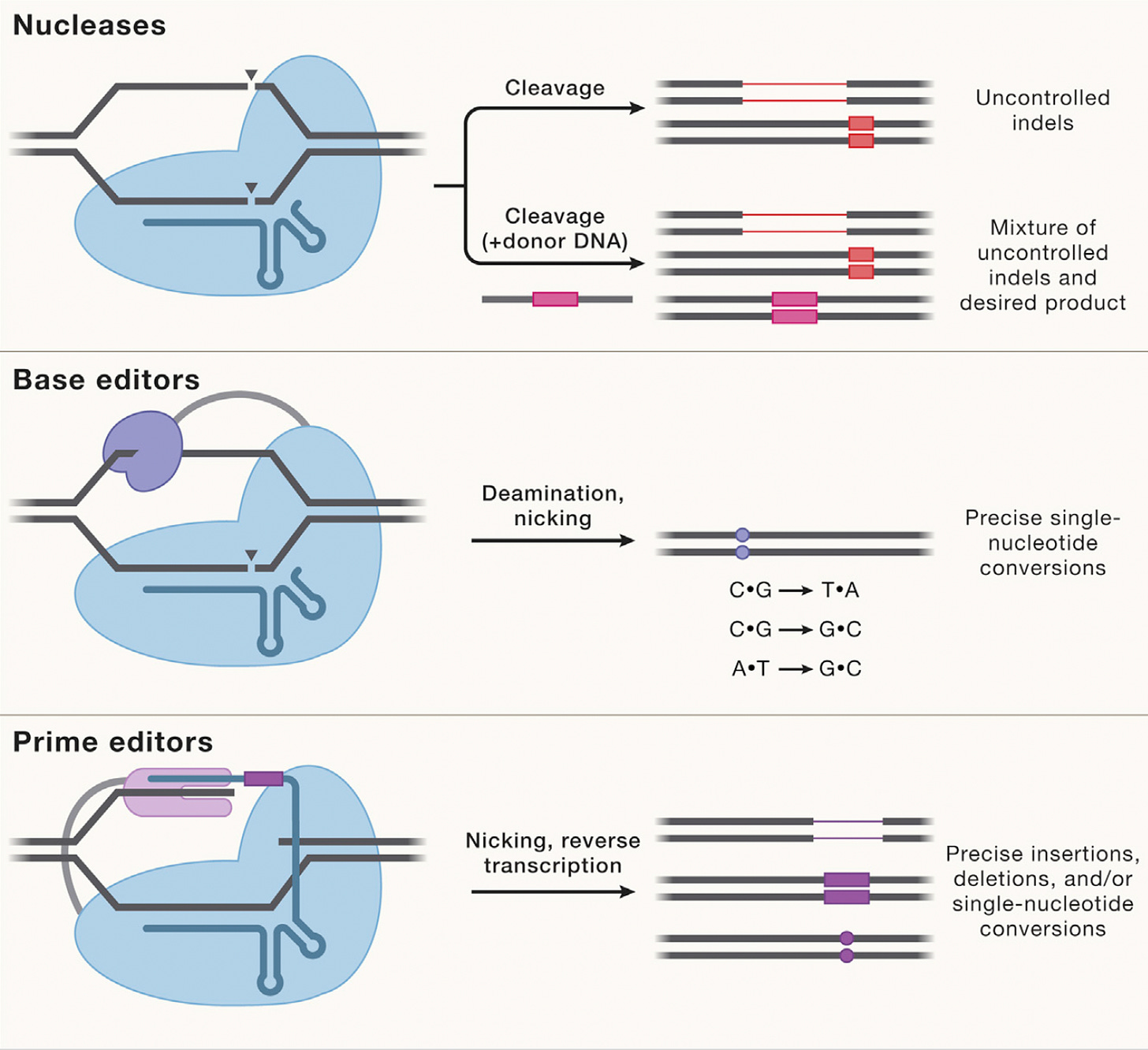
The figure below, adapted from Wang and Doudna’s Science review of the first decade of CRISPR illustrates key differences in the editing function of the 3 different modes
The Advance for Hypertrophic Cardiomyopathy
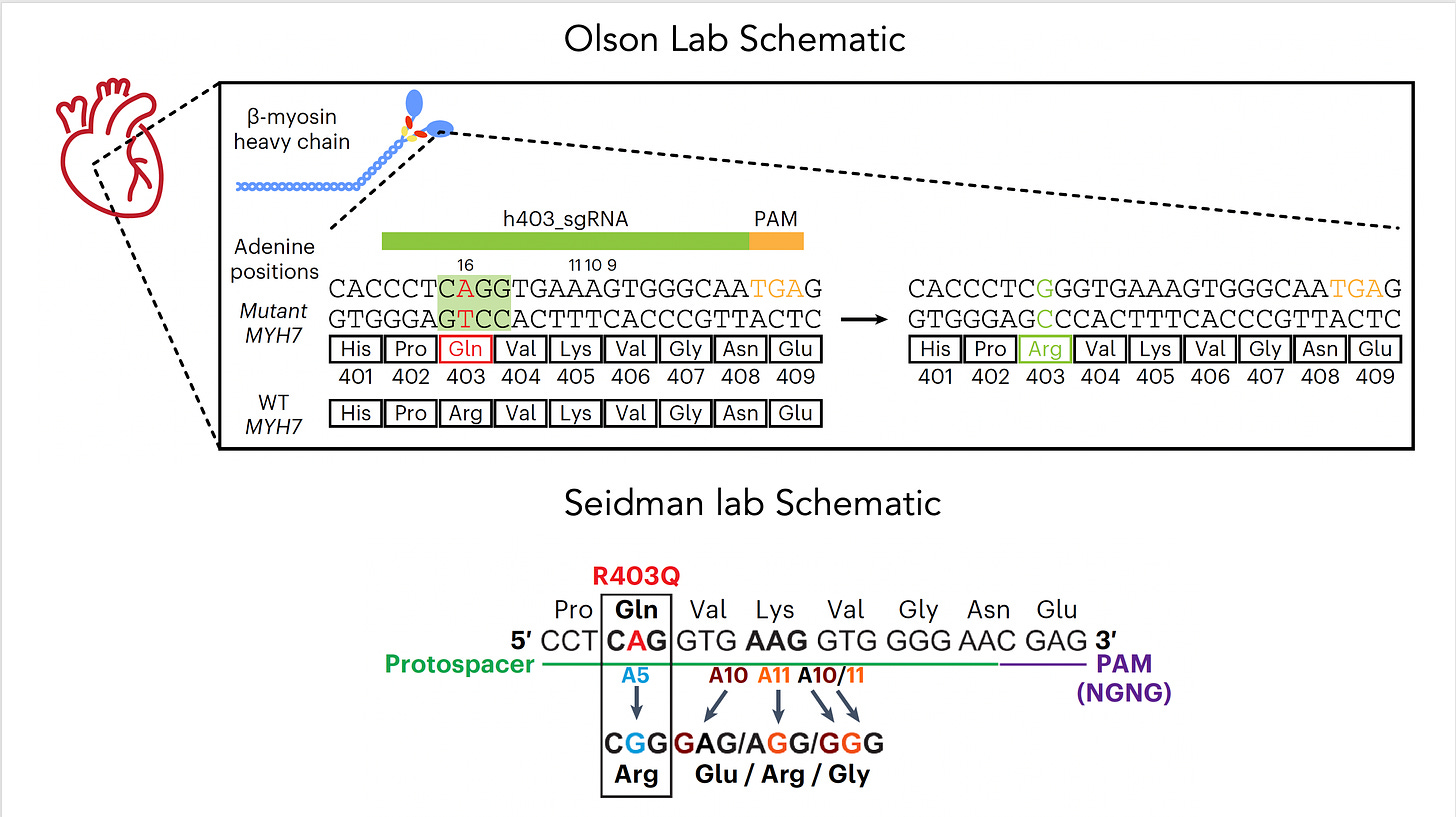
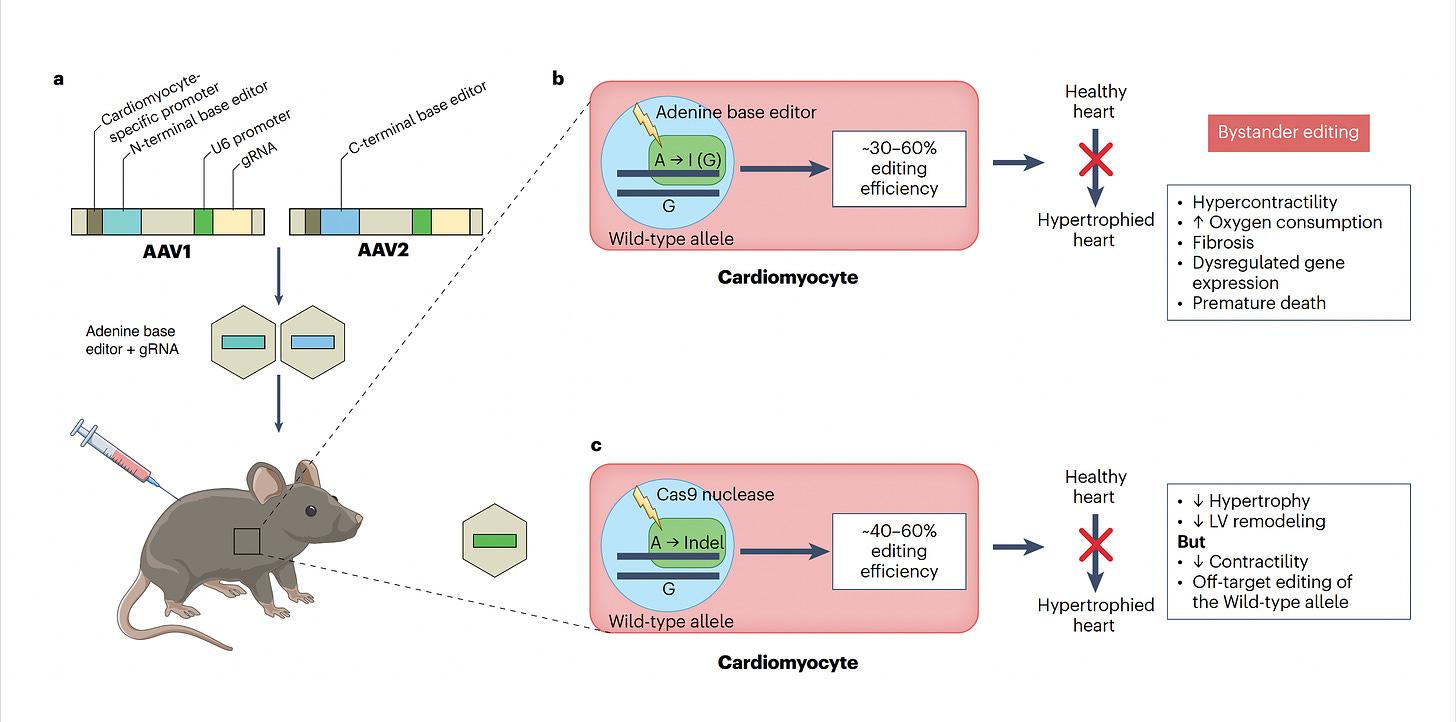
Now let’s get back to the new work on hypertrophic cardiomyopathy which was potentiated by base editing because it targeted the single nucleotide mutation—a missense in MYH7, in heart muscle’s beta-myosin, R403Q—the A (resulting in Glutamate) that needs to be fixed to a G (to get Arginine, the wild-type), perfectly suited by an adenine base editor (ABE). For some abbreviations in the schematic below, PAM stands for protospacer adjacent motif used by the base editor to find the correct site, gRNA stands for a guide RNA to promote on-target editing.
The two papers are nicely summarized in the text of the accompanying editorial and with the illustration below. To approach mouse models of HCM, both labs used base editing with an adeno-associated virus vector (AAV) delivery (discussed subsequently), a heart specific troponin promoter, with direct injection into the chest, as shown in Panel a. Editing efficiency was between 30-60% of heart muscle cells (Panel b). The Seidman lab also used Cas9 nuclease editing as shown in Panel c. There was marked amelioration of left ventricular hypertrophy, heart remodeling, and other key endpoints of rescuing or preventing heart malfunction.
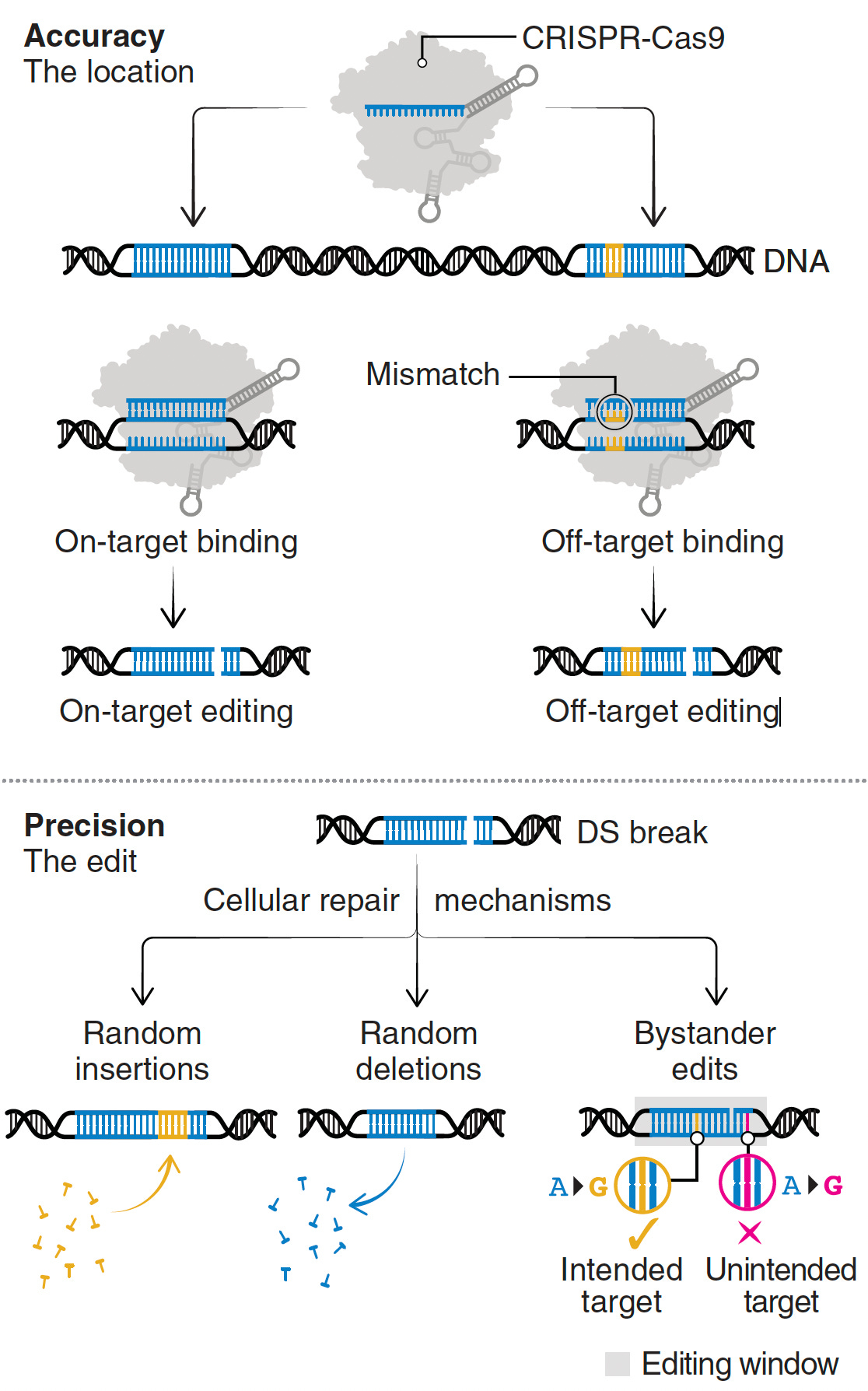
Despite these achievements, there are still multiple concerns such as bystander and off-target effects (not seen in the heterozygous model of Olson lab experiments). Bystander base editing effects refer to unintended change of bases which are still relatively common, concerning for potentially introducing pathogenic variants, but may be avoided by narrowing the editing window. With nucleases, there is a wide range of potential off-target impact that includes unintended point mutations, deletions, insertions inversions, and translocations. These unintended effects (schematic below from Wang and Doudna) are the basis of suboptimal accuracy or precision and the subject of many ongoing refinements.
Besides the unintended effects at the genome level, there is the potential of tissue “mosaicism,” whereby some cells have been fixed and others have not, setting up possible electrical dispersion and concern about arrhythmia potential. We don’t know the threshold of editing efficiency (e.g. is 30% adequate?) in people to rescue or prevent HCM. The work done to date is for a specific and important causal mutation, but there are many others that can induce HCM which would require specific guide RNAs and might not be well suited for base editing.
Imminent Refinements
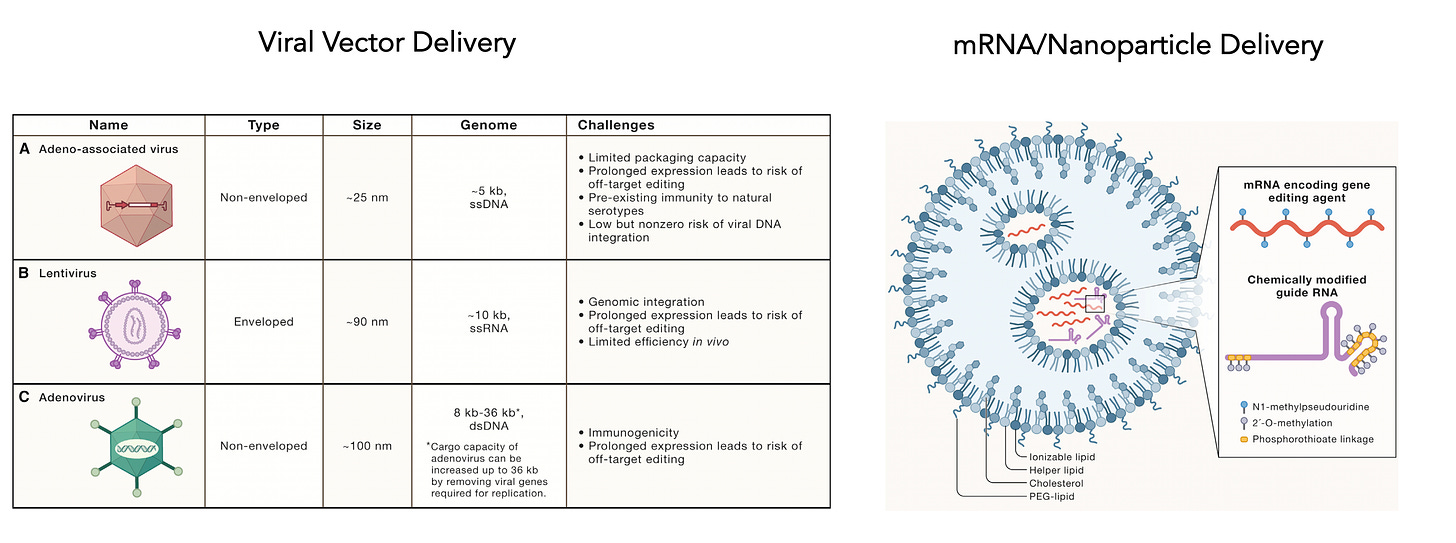
I mentioned delivery is a big issue and today there are many choices as shown below, including different viral vectors or mRNA/nanoparticle platform.
This week another pioneer in genome editing, Feng Zhang, announced a new company (Aera) with a protein-centric delivery technology called SEND which stands for Selective Endogenous eNcapsidation for cellular Delivery and may rev up efficiency of delivery. This continues to be a formidable challenge when you think of the obstacle course the genome editing package has to go through, as shown here.
Then back to precision and accuracy, which brings me to this week’s report of using AI to promote that for prime editing. The convergence of AI tools to make genome editing hit its target, avoid unintended effects, has been amplified by many papers in the past couple of years. The conclusion of the new report reinforces the power of AI convergence for improving prime editors: “Our work adds the important dimension of short sequence insertion in different DNA repair contexts, which holds promise in enabling both sophisticated genome engineering and the correction of thousands of pathogenic mutations.”
I’ve over-simplified much of this to help readers who are not initiated with all of the genome editing jargon, and even that is a difficult task. Many recent papers do not even provide what abbreviations stand for such as HNH or PAMs, assuming readers are well versed in this complicated space.
But the bottom line: watch this space. It is taking off much faster and broader than anticipated. What took 10 years to get to this point may be greatly compressed in the next few years. We’ve already seen clinical trials begin with targeting the PCSK9 gene in the liver to achieve very low LDL cholesterol to prevent heart disease, atherosclerosis, and heart attacks. That’s an easier target than going into the heart itself. But we’re on the cusp of being able to do that in the future, no less many other organs and tissues throughout the body. It’s scientifically enthralling and may (likely someday) offer cures for what previously were incurable conditions.
Thanks for reading Ground Truths. I hope you find them useful. Please subscribe to get these essays directly, as notifications via twitter and other social media platforms are unreliable (and I will likely be reducing my participation in them).